This would allow the catalytic domain to have access to pentapeptides at different distances from the membrane. It suggests that the stem domains interact and the catalytic domains have freedom to move from the position observed in the crystal structure. Weiss DS, Chen JC, Ghigo JM, Boyd D, Beckwith J: Localization of FtsI (PBP3) to the septal ring requires its membrane anchor, the Z ring, FtsA, FtsQ, and FtsL. Only 34.95 I have an idea for a project, but it needs some way to get input to the microcontroller and some way to display information from the micro.
#PBP3 DIMER SERIAL#
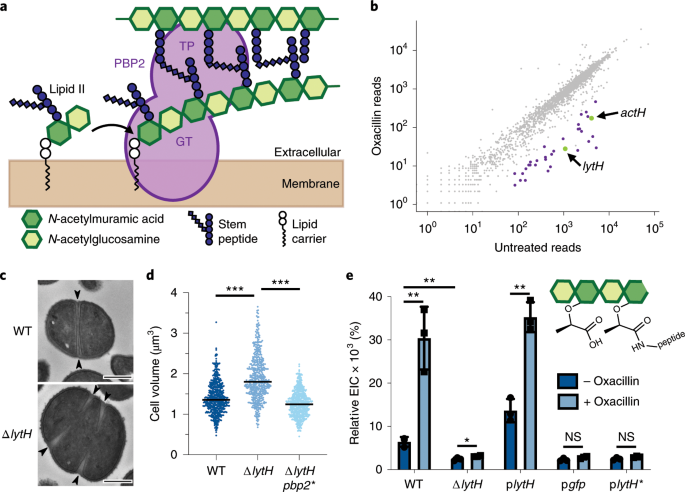
This model extends our understanding of PBP5 function as it suggests how PBP5 can interact with the peptidoglycan layer. 4-Digit LED Displays with simple, single-wire serial input. As the crystal structure of the soluble domain of PBP5 (i.e., lacking the membrane anchor) shows a monomer, we used our experimental data to generate a model of the homo‐dimer. Both approaches indicate that PBP5 exists as a homo‐oligomeric complex, most likely as a homo‐dimer. In this study, we have analyzed the oligomeric state of PBP5 in detergent and in its native environment, the inner membrane. In doing so, it varies the substrates for transpeptidation and plays a key role in maintaining cell shape. Penicillin‐binding protein 5 (PBP5) is a DD‐carboxypeptidase, which cleaves the terminal D‐alanine from the muramyl pentapeptide in the peptidoglycan layer of Escherichia coli and other bacteria. Skoog, Karl Bruzell, Filippa Stenberg Ducroux, Aurélie Hellberg, Mårten Johansson, Henrik Lehtiö, Janne Högbom, Martin Daley, Daniel O. The monofunctional class B PBP2 and PBP3 exhibit essential D,D-TPase activities associated with elongation and division, respectively (5). pneumoniae 24 but substantially extended in the present SA-PBP3 structure by five extra -helices. 29 It is slightly larger in PBP2b from S. Com-plementation studies and changes in localization suggest that these D,D-CPases are not. In PA-PBP3, this contiguous and compact domain extends over residues 80149, 19 includes four -helices and three -strands, and corresponds to the so-called PBP-dimer domain identified by Pfam. Crystal structure of penicillin-binding protein 3 (PBP3) from methicilin-resistant Staphylococcus aureus in the apo form. Penicillin‐binding protein 5 can form a homo‐oligomeric complex in the inner membrane of Escherichia coli Penicillin‐binding protein 5 can form a homo‐oligomeric complex in the inner membrane of. of Escherichia coli change dimer conformation between resting and active states.
